Featured Products
Our Promise to You
Guaranteed product quality, expert customer support

ONLINE INQUIRY

- Specification
- Background
- Scientific Data
- Q & A
- Customer Review
The GP5d cell line originates from a female patient with colon adenocarcinoma, diagnosed as locally recurrent poorly differentiated Dukes' B2 stage colon cancer at the time of surgical excision. The patient had no history of preoperative chemotherapy or radiotherapy and had a significant family history of colon cancer. The GP5d cell line exhibits epithelial-like morphology and grows in an adherent pattern. It shows excellent proliferation ability during culture but significantly differs morphologically from the GP2d cell line. GP2d cells form a uniform cell layer through diffuse growth, whereas GP5d cells form multilayered structures in discrete cell islands.
The GP5d cell line is used to study gene expression regulatory mechanisms. For example, it responds to EGF, TGF alpha, or insulin, while these factors have almost no effect on the GP2d cell line. Additionally, GP5d plays a crucial role in the study of transposable elements (TEs) derepression. Research shows that DNMTi-HDACi treatment leads to the derepression of specific TE subfamilies in GP5d, related to p53 function. GP5d cells are utilized to establish tumor models for studying cancer progression and therapeutic strategies. For instance, using CRISPR-Cas9 to delete the MYC super-enhancer region and immunoglobulin heavy chain gene (IGH) locus in GP5d allows exploration of these genes' roles in tumor development. The GP5d cell line is also used in drug screening and mechanistic studies. It was found to be most sensitive to the RRSP-1 drug, providing important insights for developing new anti-cancer therapies.
RALB is Required for the Survival of KRASMT CRC Cells
RAS gene mutations are common in CRC, resulting in poor survival and challenging treatment due to activated downstream pathways. RALA and RALB, RAS-like proteins, have roles in tumorigenesis, but their specific functions in apoptosis, especially for KRASMT CRC, remain unclear. Khawaja et al. aimed to unravel the role of RALB GTPase in apoptosis regulation, particularly its interaction with TRAIL Death Receptor 5 (DR5) in KRASMT CRC.
Khawaja's lab previously showed that RALA, not RALB, influences the migration of KRASMT CRC cells. They investigated if RALA and RALB have similar roles in cell survival (Fig. 1A). Silencing RALA slightly reduced cell viability, while siRALB significantly affected it, corroborated by additional siRNA sequences against RALB (Fig. S1A). Overexpressing WT-RALB or active G23V-RALB notably enhanced colony formation in KRASMT CRC cells (Fig. 1A). siRALB induced cell death only in KRASMT cells, evident by PARP cleavage and Caspase-3/7 activation (Fig. 1B, C; Fig. S1A–C), unlike siRALA (Fig. 1B, C; Fig. S1D). KRASMT cancers resist MEK inhibition (MEKi). They hypothesized RALB might influence MEKi resistance in KRASMT CRC. Combining RALB siRNA with MEKi AZD6244 increased cell death in HCT116 cells, shown by PARP cleavage and Caspase activity (Fig. 1B; Fig. S1B), but not in KRASMT cells (Fig. 1B; Fig. S1C). In KRASMT cell lines (SW620, GP5D, LoVo), AZD6244 added to siRALB didn't increase cell death beyond siRALB alone (Fig. 1C; Fig. S1D). AZD6244 with siRALA showed no effect. No changes in pERK1/2 levels were observed. Hence, RALB, not RALA, regulates survival in KRASMT but not KRASMT CRC cells.
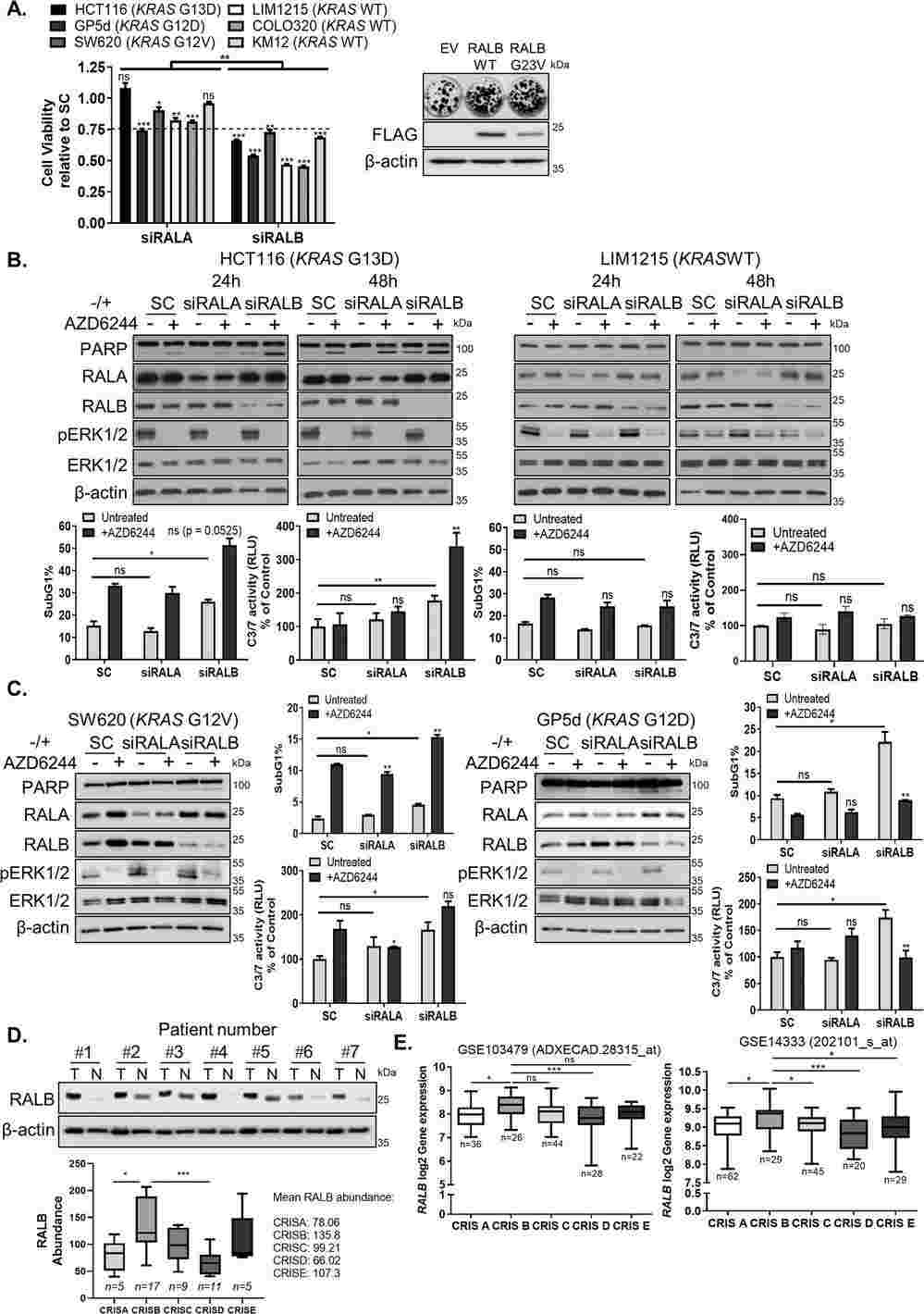 Fig. 1. RASMT CRC cells are dependent on RALB for survival (Khawaja H, Campbell A, et al., 2020).
Fig. 1. RASMT CRC cells are dependent on RALB for survival (Khawaja H, Campbell A, et al., 2020).
RRSP Inhibits Proliferation and Perk Activation in CRC Cell Lines
Ras oncoproteins, such as KRAS, play a crucial role in cell signaling related to proliferation and survival. Mutations in RAS genes are present in around 30% of human cancers and significantly drive tumorigenesis, making RAS a prime target for cancer therapy.
To examine the impact of RAS processing on downstream signaling, Stubbs et al. analyzed four KRAS mutant CRC cell lines (HCT-116, SW1463, SW620, GP5d) with different KRAS mutations (Fig. 2A). Due to varying expression of the anthrax toxin receptor, they used a potent RRSP chimeric toxin (RRSP-DTB) linked to the diphtheria toxin's translocation fragment. RRSP-DTB binds to the human HB-EGF receptor, is endocytosed, and translocates into the cytosol. The HB-EGF receptor expression was consistent across CRC cell lines. They treated cells with increasing doses of RRSP-DTB or inactive RRSP*-DTB, monitoring growth inhibition. HCT-116 cells were most affected, showing the lowest IC50 (Fig. 2B), confirming high susceptibility to RRSP. Inactive RRSP*-DTB had no effect, verifying that RAS processing caused the sensitivity (Fig. 2B). SW1463 cells were also sensitive but had a 12-fold higher IC50 than HCT-116 (Fig. 2C). SW620 showed about 40% growth inhibition with RRSP-DTB at 10 nM, reflecting prior categorization of growth inhibition response (Fig. 2D, E). GP5d was similarly less susceptible but showed some growth inhibition. RRSP cleaved at least 80% of RAS across all lines and reduced ERK phosphorylation compared to RRSP*-DTB samples (Fig. 2F-H). There was variability in uncleaved RAS detection, likely due to compensatory RAS upregulation, especially in HCT-116 (Fig. 2F).
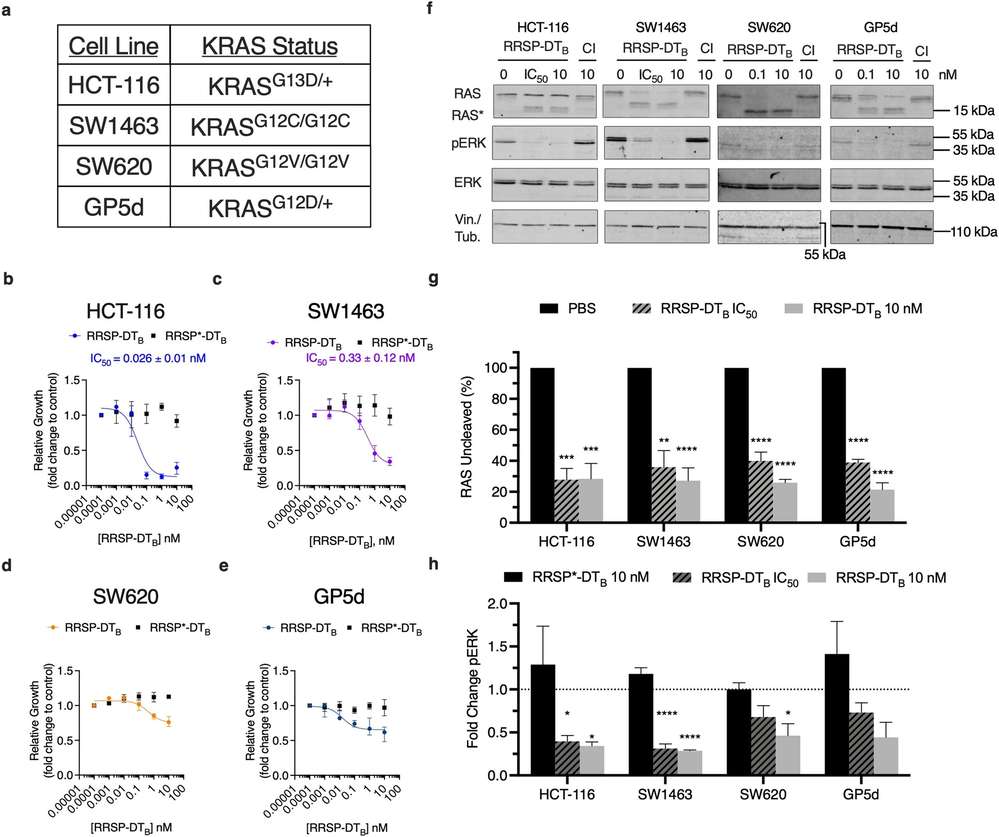 Fig. 2. RRSP-DTB growth inhibition in CRC cell lines (Stubbs CK, Biancucci M, et al., 2021).
Fig. 2. RRSP-DTB growth inhibition in CRC cell lines (Stubbs CK, Biancucci M, et al., 2021).
The process of restoring cell growth by thawing cells that have been frozen in liquid nitrogen or -70°C and then re-cultured. When restored to a normothermic state, the morphological structure of the cells remains normal and biochemical reactions can be restored.
Ask a Question
Average Rating: 5.0 | 1 Scientist has reviewed this product
Satisfied
I'm very satisfied with the services and products of various cell, and I look forward to working with them again in the future.
14 Mar 2022
Ease of use
After sales services
Value for money
Write your own review